Deep Learning and EEG
A growing part of my research applies deep learning to large-scale EEG datasets. These projects address practical obstacles that have limited the use of neural networks in EEG research -- notably the heterogeneity of electrode montages across laboratories and the question of whether raw time-domain signals or engineered spectral features provide better input representations. The work spans classification architectures, channel adaptation strategies, spatial attention mechanisms, and systematic benchmarking of pretrained foundation models.
Raw EEG vs spectral features for deep learning
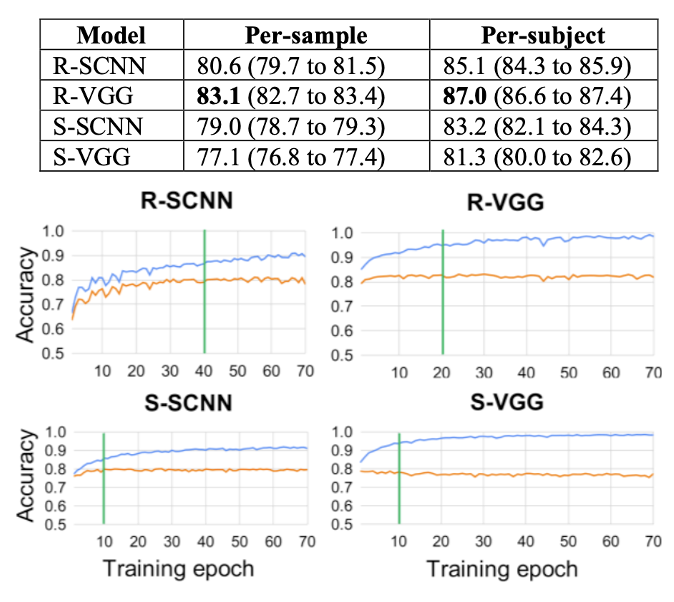
This study compared convolutional neural networks trained on raw EEG time series against networks trained on spectral topographic features for the task of biological sex classification. Using 128-channel resting-state recordings from 1,574 participants in the Healthy Brain Network, we found that a simple CNN operating on raw 24-channel × 256-sample matrices outperformed the more complex spectral-topography approach (VGG-16 adapted for EEG). The result suggested that hand-crafted spectral features, while interpretable, may discard temporal information that raw-data models can exploit. This was one of the first large-cohort comparisons of input representations for EEG deep learning.
Truong D, Milham M, Makeig S, Delorme A. Deep convolutional neural network applied to electroencephalography: Raw Data vs spectral features. IEEE EMBC, 2021. [arXiv]
Riemannian recentering for channel adaptation
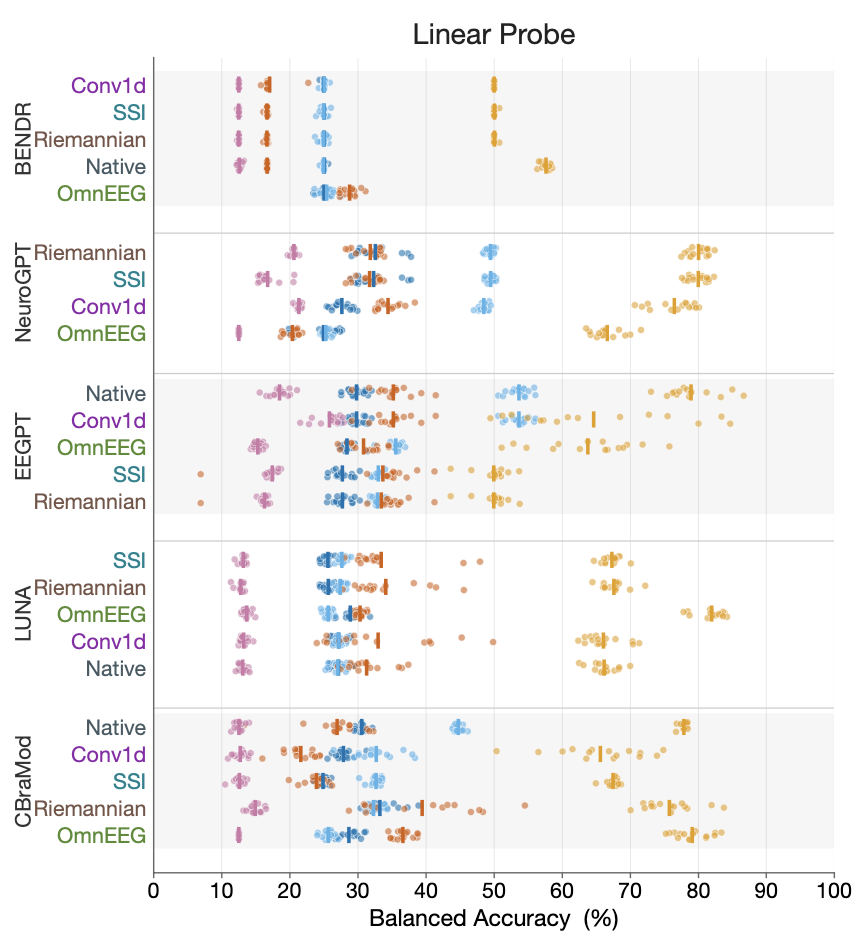
Training deep learning models across EEG datasets is difficult because electrode montages differ between laboratories. This benchmark evaluated four channel-adaptation strategies -- zero-padding, spherical interpolation, learned channel tokens, and Riemannian recentering -- across five pretrained EEG foundation models. Riemannian recentering, a fixed geometric operation on the symmetric positive definite manifold of covariance matrices that requires no learned parameters, performed on par with the best learned adaptation methods. The finding implies that a simple preprocessing step can replace expensive retraining when transferring pretrained models to new montages.
Kokate A, Aristimunha B, Truong D, Delorme A. Riemannian recentering for EEG foundation models. arXiv, 2026.
Spatial attention for cross-montage learning
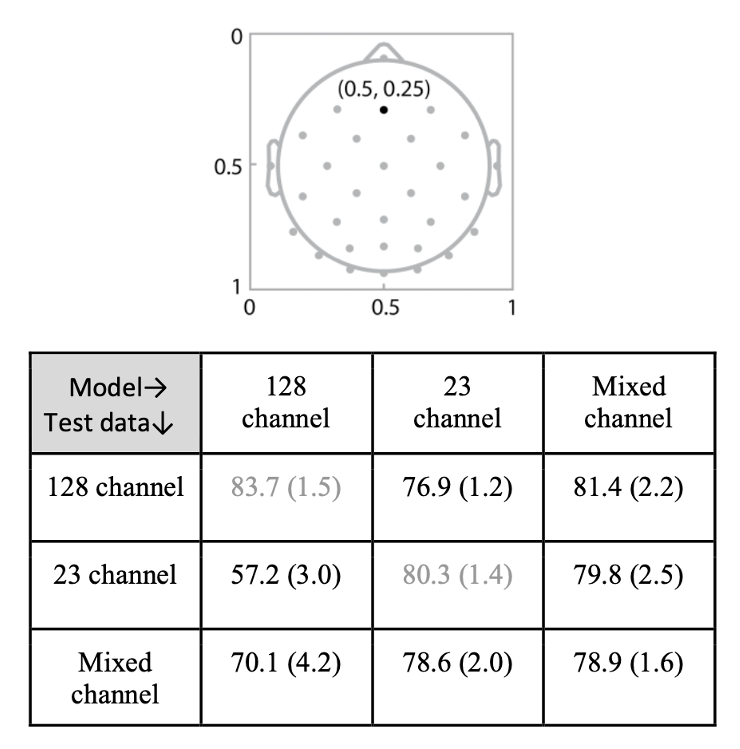
Different EEG laboratories use different electrode montages, making it difficult to combine datasets for deep learning. This paper introduced a spatial attention mechanism based on the Discrete Fourier Transform that learns a continuous mapping from electrode coordinates to attention weights. The mechanism is independent of the number and location of input channels: it transforms raw electrode positions into a 2-D frequency space, enabling the network to generalize across montages without retraining. Experiments on gender classification showed that models trained on 128-channel data with spatial attention maintained performance when tested on 23-channel montages, and that mixed-montage training further improved results.
Truong D, Makeig S, Delorme A. Deep learning applied to EEG data with different montages using spatial attention. IEEE BIBM, 2023. [arXiv]
OpenEEG-Bench: benchmarking EEG foundation models
OpenEEG-Bench is a systematic benchmark for evaluating pretrained EEG foundation models across standardized tasks. The project provides a unified evaluation protocol with a live leaderboard on HuggingFace, allowing researchers to compare model performance on classification and regression tasks drawn from multiple public datasets. By fixing the evaluation framework, OpenEEG-Bench aims to bring the kind of reproducible comparison common in computer vision and NLP to the EEG deep learning community.
Guetschel P, Aristimunha B, Truong D, Kokate A, Tangermann, M, Delorme AOpenEEG-Bench. Submitted to EUSIPCO, 2026.